Rakesh David1,2, R. Daniel Kortschak1, and Iain Searle1§
1School of Biological Sciences, The University of Adelaide, The University of Adelaide and Shanghai Jiao Tong University Joint International Centre for Agriculture and Health, Adelaide, South Australia 5005, Australia
2Present address: ARC Centre of Excellence in Plant Energy Biology, The School of Agriculture, Food and Wine, University of Adelaide, Waite Campus, Adelaide, South Australia 5005, Australia
§Correspondence to: Iain Searle (iain.searle@adelaide.edu.au)
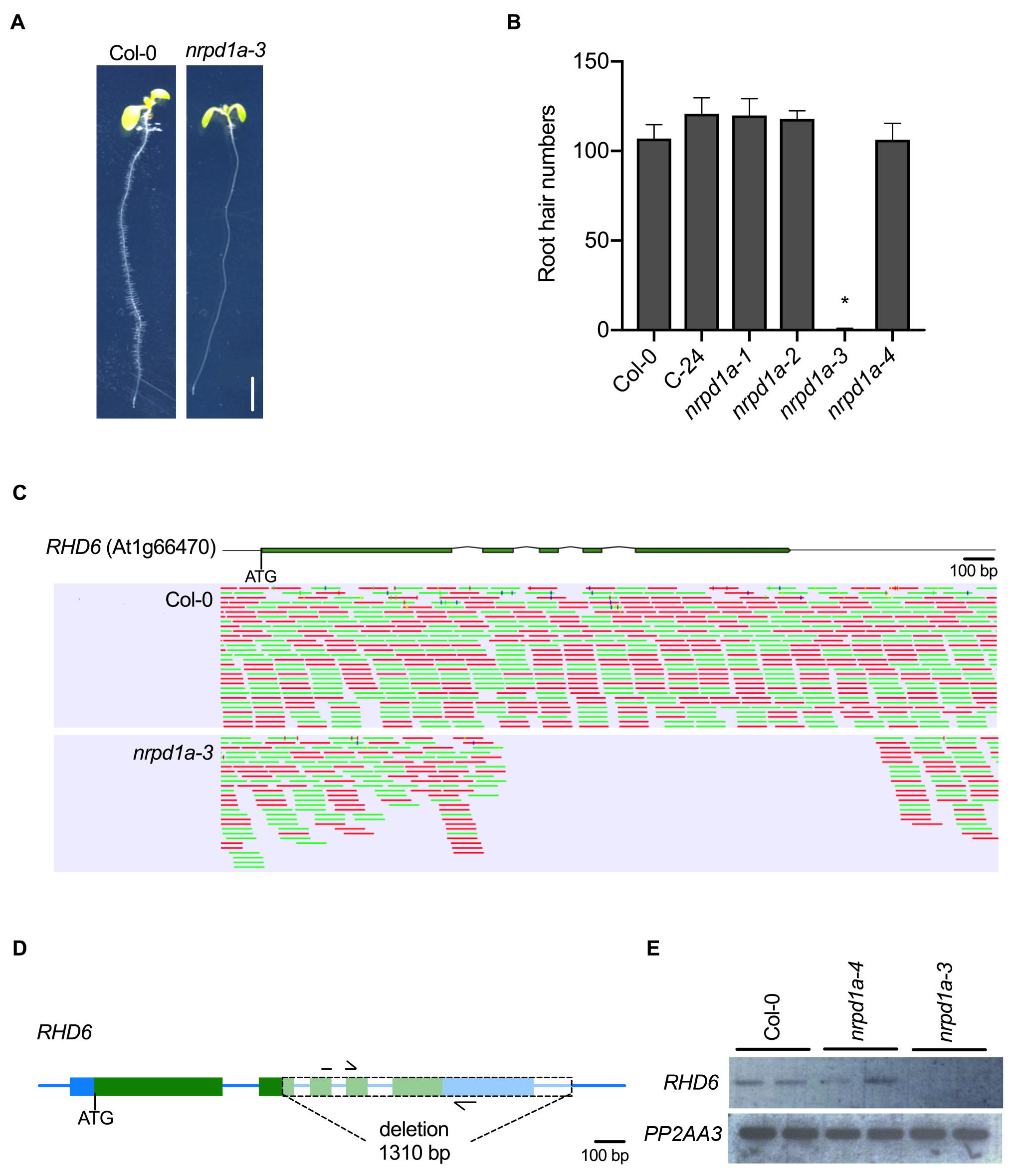
Description
Reagents
Acknowledgements
Funding
Author Contributions
- Rakesh David: Investigation, Writing - original draft, Writing - review and editing, Visualization
- R. Daniel Kortschak: Investigation, Writing - review and editing
- Iain Searle: Investigation, Writing - original draft, Writing - review and editing, Visualization, Conceptualization, Funding acquisition, Project administration, Supervision
Reviewed By
Anonymous
History
- Received: 11/20/2019
- Accepted: 12/5/2019
- Published: 12/12/2019
Copyright
© 2019 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: 7252326
PubMed: 32550444

