Aimee Jaramillo-Lambert1,2§, and Andy Golden1
1National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892
2Presently-Department of Biological Sciences, University of Delaware, Newark, Delaware 19701
§Correspondence to: Aimee Jaramillo-Lambert (anjl@udel.edu)
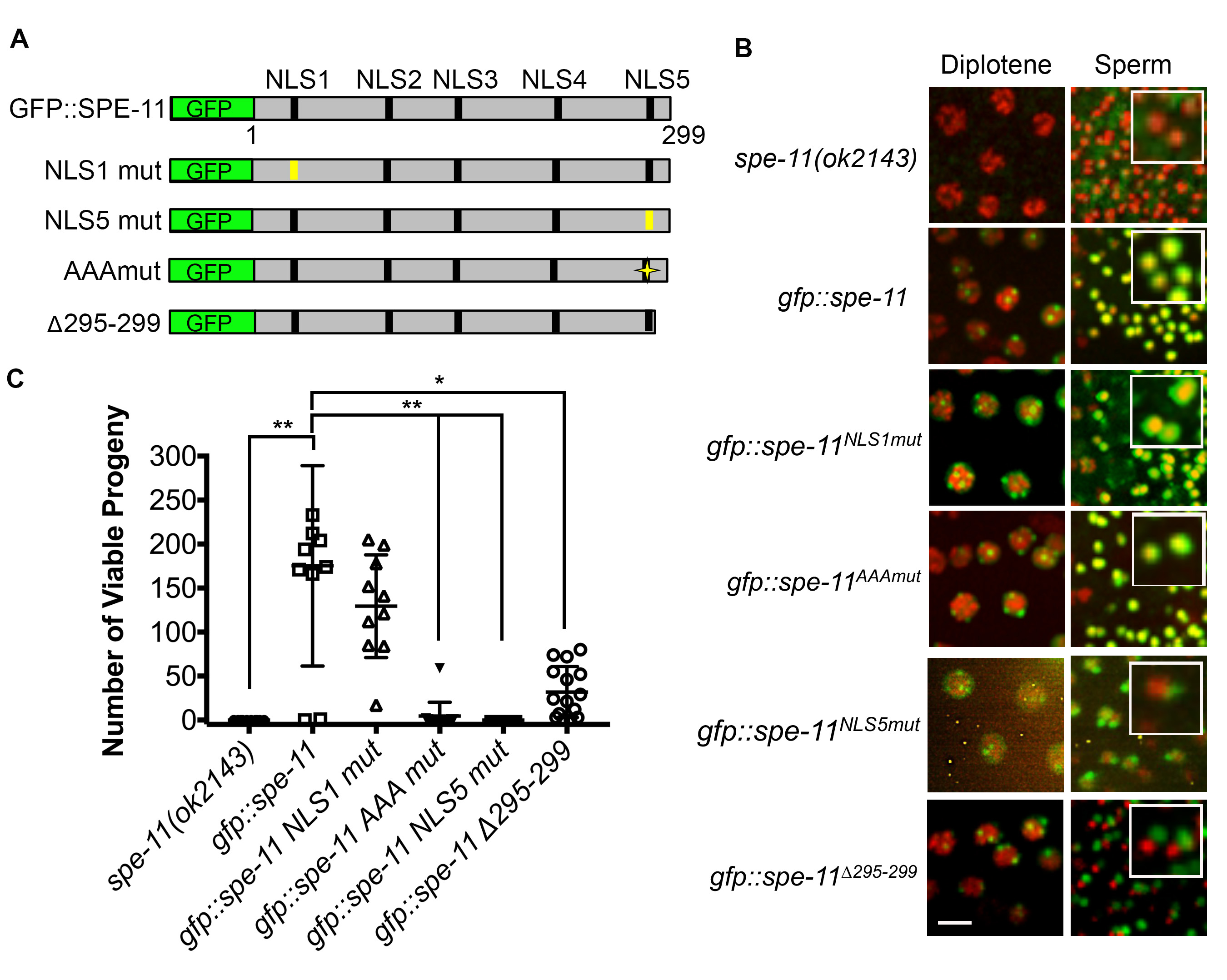
Description
Methods
Reagents
Acknowledgements
Funding
Author Contributions
- Aimee Jaramillo-Lambert: Formal analysis, Investigation, Conceptualization, Writing - original draft, Writing - review and editing
- Andy Golden: Conceptualization, Supervision, Writing - review and editing
Reviewed By
Susan Strome
Database Reference ID: WBPaper00059655
History
- Received: 5/12/2020
- Revision Received: 5/19/2020
- Accepted: 5/19/2020
- Published: 5/21/2020
Copyright
© 2020 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC7252397
PubMed: 32550511
microPublication Biology:ISSN: 2578-9430

