Bhagyashree Kaduskar1, Deepti Trivedi2, and Girish S Ratnaparkhi1§
1Indian Institute of Science Education & Research, Pune, 411008 INDIA
2Fly Facility, National Centre for Biological Sciences (NCBS), TIFR, Bangalore 560065 INDIA
§Correspondence to: Girish S Ratnaparkhi (girish@iiserpune.ac.in)
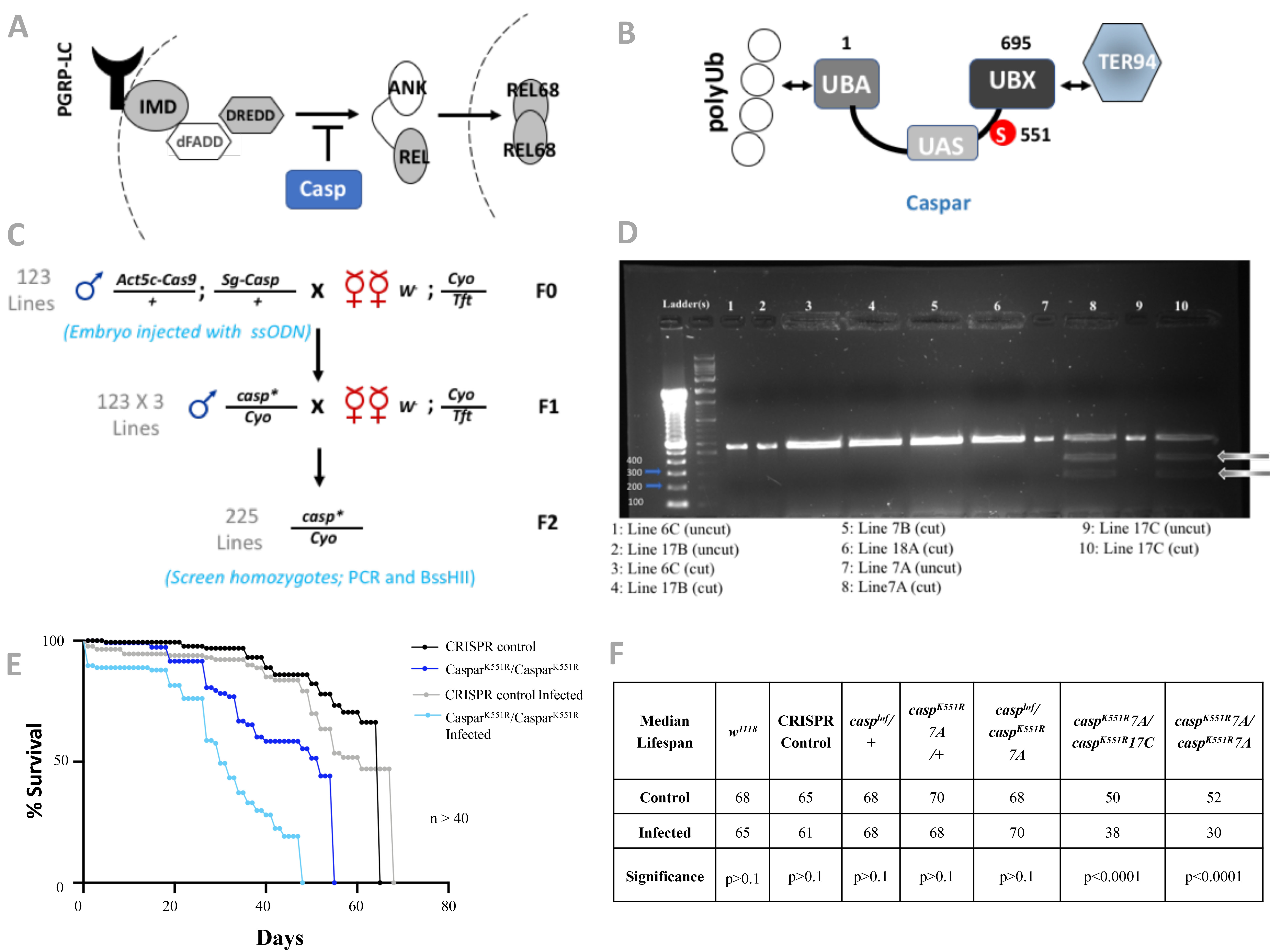
Description
Methods
Funding
Author Contributions
- Bhagyashree Kaduskar: Conceptualization, Investigation, Writing - original draft, Visualization, Formal analysis
- Deepti Trivedi: Investigation, Methodology, Writing - review and editing, Conceptualization, Validation, Data curation, Formal analysis
- Girish S Ratnaparkhi: Supervision, Writing - original draft, Conceptualization, Methodology, Writing - review and editing, Formal analysis, Investigation
Reviewed By
Anonymous, Steven Marygold
History
- Received: 5/15/2020
- Revision Received: 7/23/2020
- Accepted: 7/23/2020
- Published: 8/2/2020
Copyright
© 2020 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: 7396161
PubMed: 32760885
microPublication Biology:ISSN: 2578-9430

