Housei Wada1§, and Shigeo Hayashi1§
1RIKEN Center for Biosystems Dynamics Research
§Correspondence to: Housei Wada (housei.wada@riken.jp) Shigeo Hayashi (shigeo.hayashi@riken.jp)
Description
Reagents
Extended Data
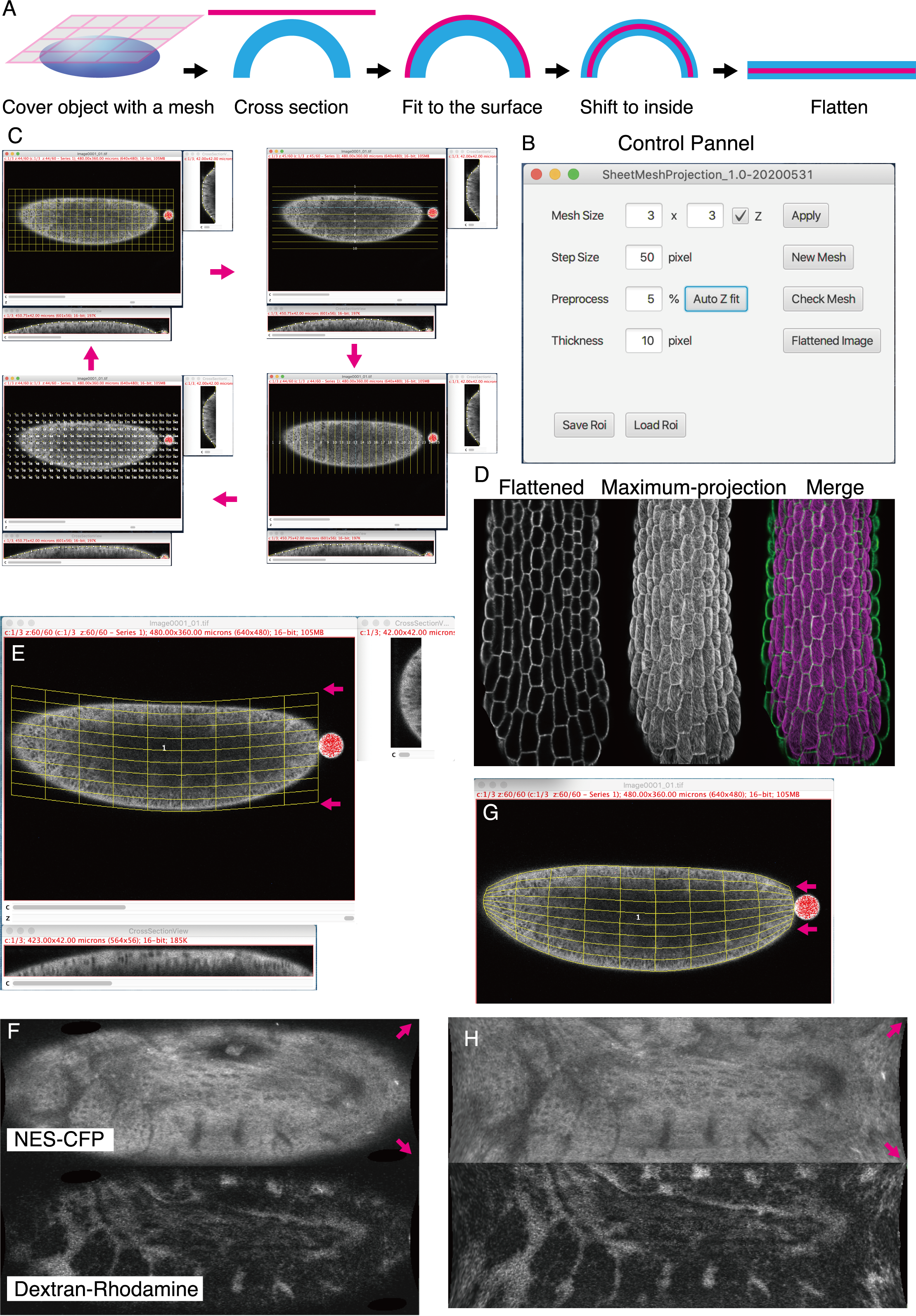
- Description: Methods: A highly versatile ImageJ plugin “SheetMeshProjection” for extracting curved surfaces with user-defined thickness and creating expanded 2D images.. Resource Type: Software. DOI: 10.22002/D1.1606
Acknowledgements
Funding
Author Contributions
- Housei Wada: Conceptualization, Formal analysis, Methodology, Software, Writing - original draft
- Shigeo Hayashi: Writing - review and editing, Project administration
Reviewed By
Steven Marygold, Pavel Tomancak
History
- Received: 4/30/2020
- Revision Received: 8/17/2020
- Accepted: 8/19/2020
- Published: 8/19/2020
Copyright
© 2020 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC7443341
PubMed: 32844154
microPublication Biology:ISSN: 2578-9430

