Mikiya Umeyama1, and Kengo Morohashi1§
1Faculty of Science and Technology, Department of Applied Biological Science, Tokyo University of Science, 2641 Yamazaki, Noda, Chiba 278-8510, Japan
§Correspondence to: Kengo Morohashi (morohashi.1@rs.tus.ac.jp)
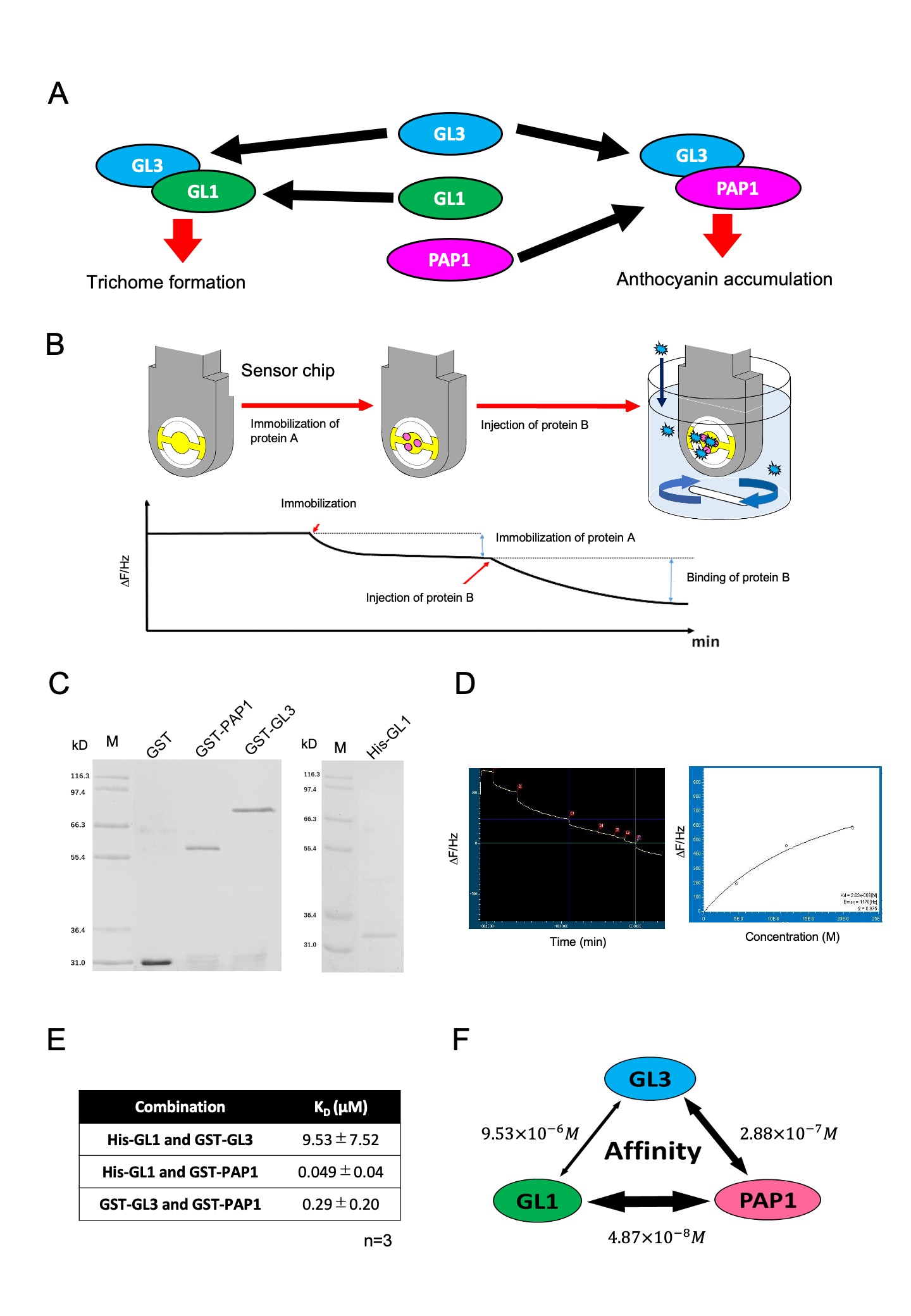
Description
Methods
Acknowledgements
Funding
Author Contributions
- Mikiya Umeyama: Investigation, Data curation, Validation
- Kengo Morohashi: Conceptualization, Data curation, Supervision, Writing - original draft, Writing - review and editing, Funding acquisition
Reviewed By
Anonymous
Nomenclature Validated By
Anonymous
History
- Received: 6/26/2020
- Revision Received: 8/17/2020
- Accepted: 8/18/2020
- Published: 8/19/2020
Copyright
© 2020 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC7443340
PubMed: 32844153
microPublication Biology:ISSN: 2578-9430

