Lucila Buzzi1*, Victoria Ayelen Segobia2*, Diego Rayes3,4*, and Olga A Castro5,6§
1Fundación Instituto Leloir, Buenos Aires, Argentina
2Departamento de Fisiología, Biología Molecular y Celular, Facultad de Ciencias Exactas y Naturales, Universidad de Buenos Aires, Buenos Aires, Argentina
3Instituto de Investigaciones Bioquímicas de Bahía Blanca (CONICET)
4 Departamento de Biología, Bioquímica y Farmacia, Universidad Nacional del Sur, Bahía Blanca, Argentina.
5Consejo Nacional de Investigaciones Científicas y Técnicas
6Instituto de Biociencias, Biotecnología y Biología traslacional, Departamento de Fisiología, Biología Molecular y Celular, Facultad de Ciencias Exactas y Naturales, Universidad de Buenos Aires, Buenos Aires, Argentina
§Correspondence to: Olga A Castro (alecastro2901@gmail.com)
* Authors have equal contribution
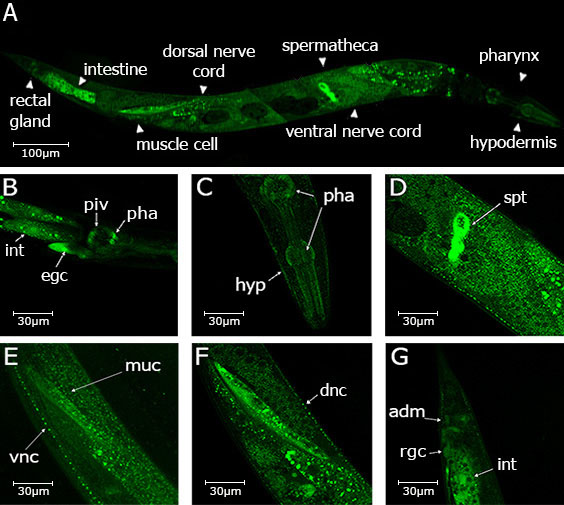
Description
Methods
Acknowledgements
Funding
Author Contributions
- Lucila Buzzi: Investigation, Conceptualization
- Victoria Ayelen Segobia: Software, Investigation
- Diego Rayes: Investigation, Methodology, Resources
- Olga A Castro: Funding acquisition, Writing - review and editing, Conceptualization, Supervision
Reviewed By
Anonymous
Database Reference ID: WBPaper00060116
History
- Received: 7/30/2020
- Revision Received: 8/20/2020
- Accepted: 8/24/2020
- Published: 8/25/2020
Copyright
© 2020 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC7456565
PubMed: 32885150

