Shalini Menon1, Dennis Goldfarb1, Emily M. Cousins1, M. Ben Major1, and Stephanie L. Gupton1§
1University of North Carolina at Chapel Hill
§Correspondence to: Stephanie L. Gupton (sgupton@unc.edu)
Description
Methods
Extended Data
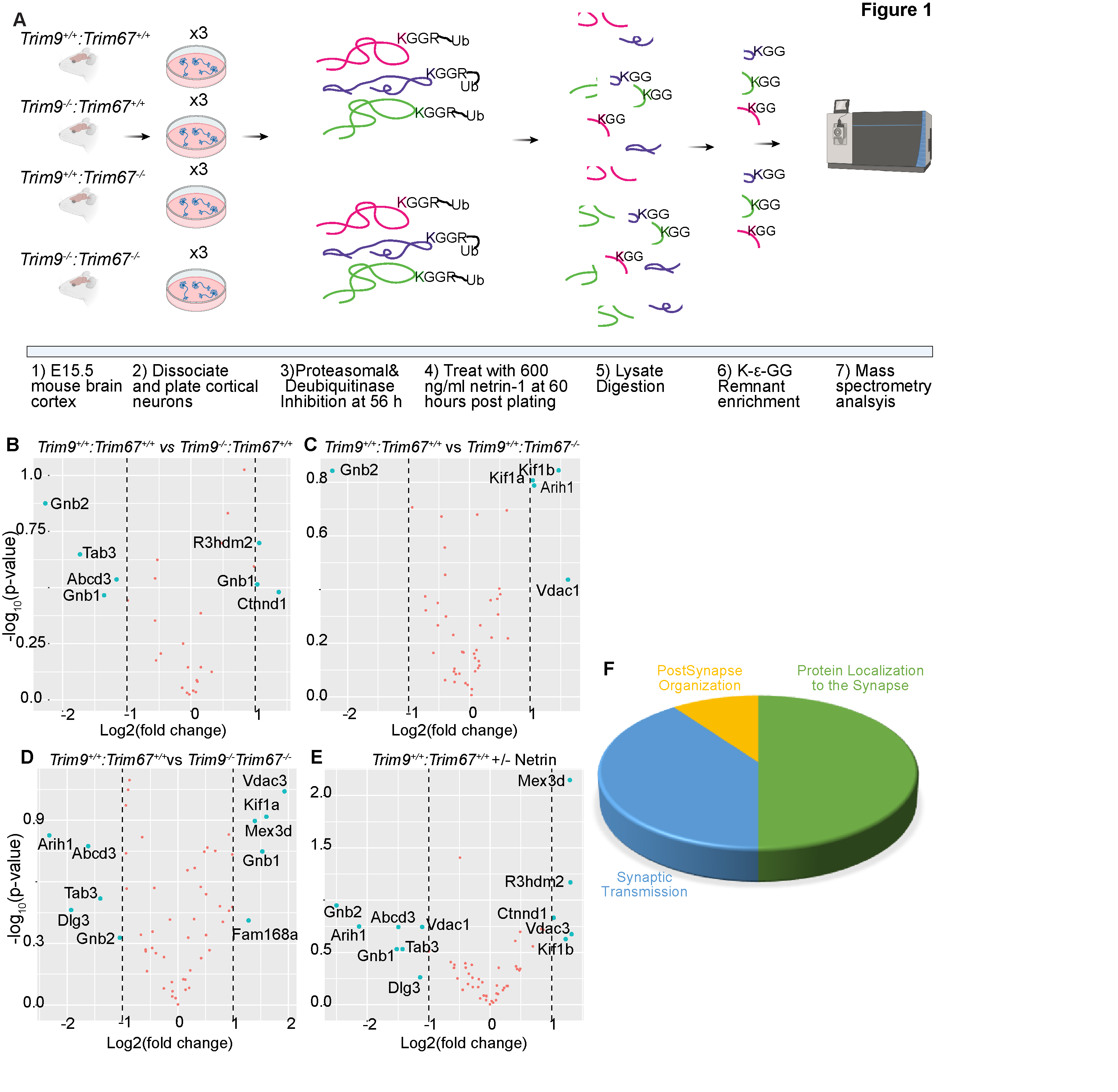
- Description: Ubiquitylome of four mouse genotypes and netrin treatment in triplicate. These data is composed of a list of 854 unique proteins and 1454 unique peptides that were ubiquitylated during early neuronal development. The data are represented in sheets that include gene names of identified proteins with NCBI identifiers; the peptide sequences identified by MS; the peptides with ubiquiylated lysine; the number of glycine adducts per peptide; the types of samples (genotype and netrin treatment) and replicates; and median subtracted intensity before and after batch correction.. Resource Type: Dataset. DOI: 10.22002/D1.1678
Acknowledgements
Funding
Author Contributions
- Shalini Menon: Conceptualization, Data curation, Formal analysis, Investigation, Writing - original draft, Writing - review and editing
- Dennis Goldfarb: Data curation, Investigation, Methodology
- Emily M. Cousins: Formal analysis
- M. Ben Major: Project administration, Writing - review and editing
- Stephanie L. Gupton: Conceptualization, Funding acquisition, Methodology, Project administration, Resources, Supervision, Writing - original draft
Reviewed By
Anonymous
History
- Received: 10/13/2020
- Revision Received: 11/11/2020
- Accepted: 11/14/2020
- Published: 11/28/2020
Copyright
© 2020 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC7704252
PubMed: 33274322
microPublication Biology:ISSN: 2578-9430

