Arnaud Ahier1,2, Shashi Kumar Suman1, and Sophie Jarriault1§
1IGBMC, Development and Stem Cells Department, CNRS UMR7104, INSERM U1258, Université de Strasbourg, Illkirch CU Strasbourg, 67404 France
2current address: Clem Jones Centre for Ageing Dementia Research, Queensland Brain Institute, The University of Queensland, Brisbane, Australia
§Correspondence to: Sophie Jarriault (sophie@igbmc.fr)
Abstract
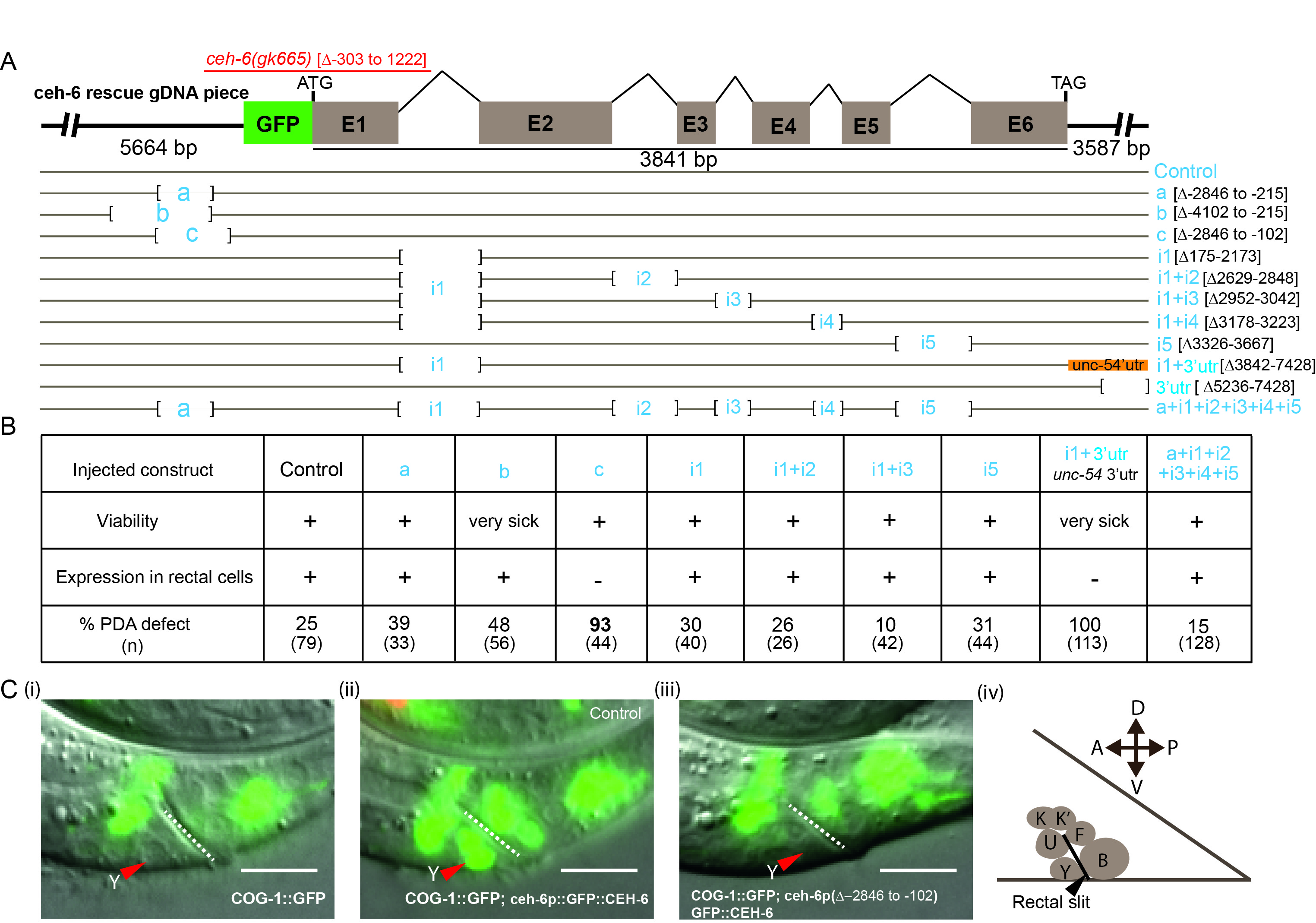
Description
Methods
Reagents
Acknowledgements
Funding
Author Contributions
- Arnaud Ahier: Conceptualization, Investigation, Methodology, Validation, Formal analysis, Writing - review and editing
- Shashi Kumar Suman: Visualization, Data curation, Formal analysis, Writing - original draft, Writing - review and editing
- Sophie Jarriault: Conceptualization, Formal analysis, Supervision, Project administration, Funding acquisition, Writing - review and editing
Reviewed By
Anonymous
Nomenclature Validated By
Anonymous
Database Reference ID: WBPaper00060684
History
- Received: 10/26/2020
- Revision Received: 12/7/2020
- Accepted: 12/15/2020
- Published: 12/21/2020
Copyright
© 2020 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC7753896
PubMed: 33364555

