Verónica A. Segarra1§, Anupam Sharma2, and Sandra K. Lemmon3
1Department of Biology, High Point University, High Point, NC, USA 27268
2Department of Microbiology, University of Georgia, Athens, GA, USA 30602
3Department of Molecular and Cellular Pharmacology, University of Miami Miller School of Medicine, Miami, FL, USA 33101
§Correspondence to: Verónica A. Segarra (vsegarra@highpoint.edu)
Abstract
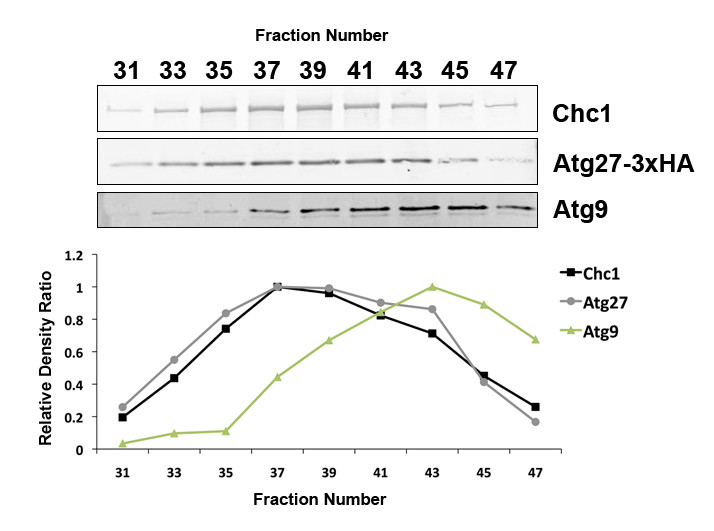
Description
Methods
Extended Data
- Description: CCVs clathrin null strain, all mass spectrometry hits, sorted, and with abbreviated information per slice.. Resource Type: Dataset. DOI: 10.22002/D1.1925
- Description: CCVs clathrin null strain, all mass spectrometry hits with full information, one slice per tab.. Resource Type: Dataset. DOI: 10.22002/D1.1926
Acknowledgements
Funding
Author Contributions
- Verónica A. Segarra: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Validation, Visualization, Writing - original draft, Writing - review and editing
- Anupam Sharma: Formal analysis, Writing - review and editing
- Sandra K. Lemmon: Conceptualization, Data curation, Funding acquisition, Methodology, Resources, Writing - review and editing, Formal analysis
Reviewed By
Anonymous
History
- Received: 1/4/2021
- Revision Received: 3/21/2021
- Accepted: 3/24/2021
- Published: 3/29/2021
Copyright
© 2021 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC8008255
PubMed: 33817564
microPublication Biology:ISSN: 2578-9430

