Hiroshi Qadota1, Andres F Oberhauser2, and Guy M Benian1§
1Department of Pathology, Emory University, Atlanta, Georgia
2Department of Neuroscience, Cell Biology & Anatomy, Sealy Center for Structural Biology and Molecular Biophysics, University of Texas Medical Branch, Galveston, Texas
§Correspondence to: Guy M Benian (pathgb@emory.edu)
Abstract
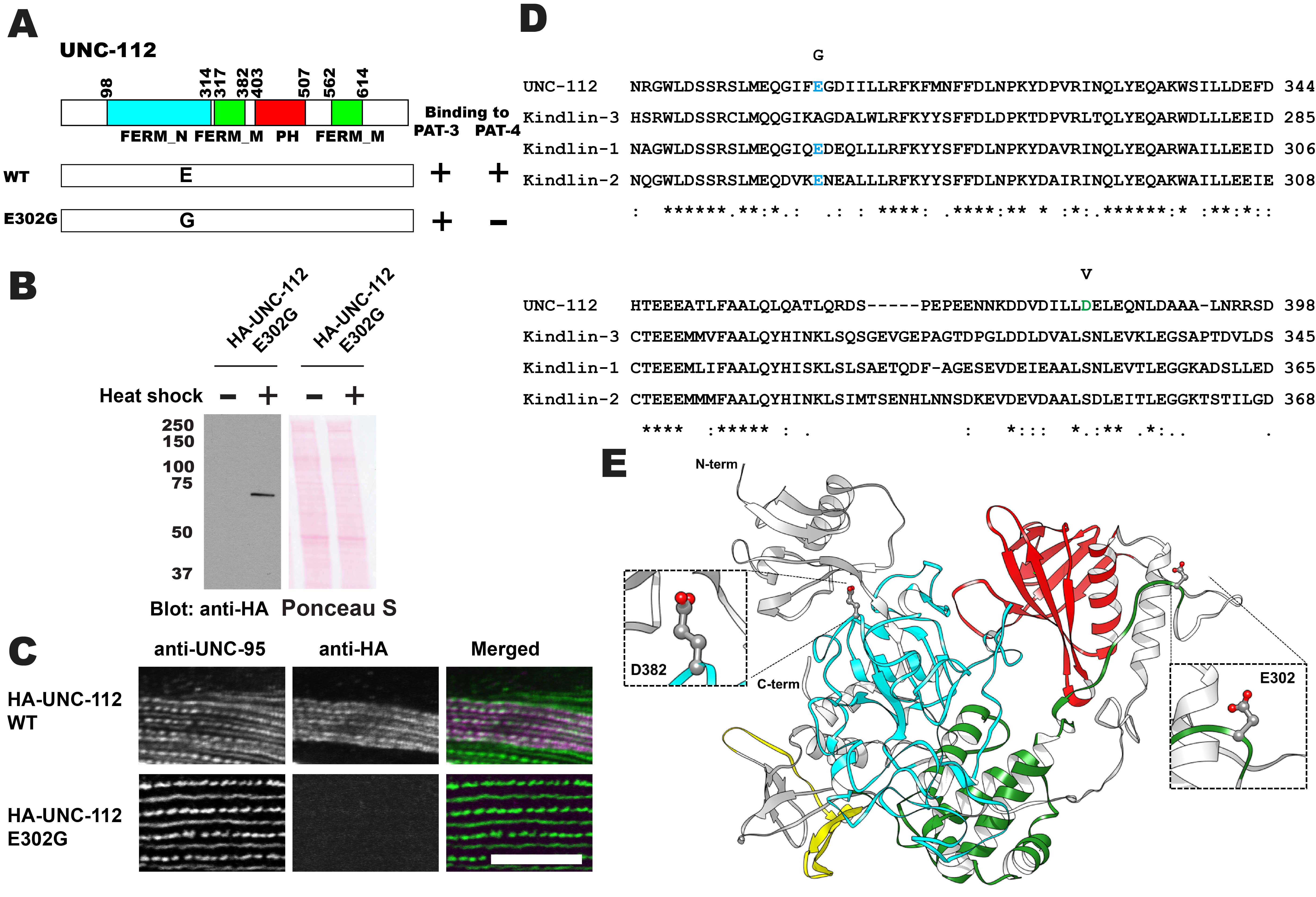
Description
Methods
Acknowledgements
Funding
Author Contributions
- Hiroshi Qadota: Conceptualization, Data curation, Investigation, Methodology, Project administration, Resources, Supervision, Validation, Visualization, Writing - original draft
- Andres F Oberhauser: Data curation, Investigation, Methodology, Validation, Writing - original draft, Writing - review and editing
- Guy M Benian: Conceptualization, Funding acquisition, Project administration, Writing - original draft, Writing - review and editing
Reviewed By
Anonymous
Nomenclature Validated By
Anonymous
Database Reference ID: WBPaper00061880
History
- Received: 8/10/2021
- Revision Received: 9/2/2021
- Accepted: 9/7/2021
- Published: 9/14/2021
Copyright
© 2021 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC8449257
PubMed: 34549173
microPublication Biology:ISSN: 2578-9430

