Michael Nonet
Dept of Neuroscience, Washington University Medical School, St. Louis MO 63110
Correspondence to: Michael Nonet nonetm@wustl.edu
Abstract
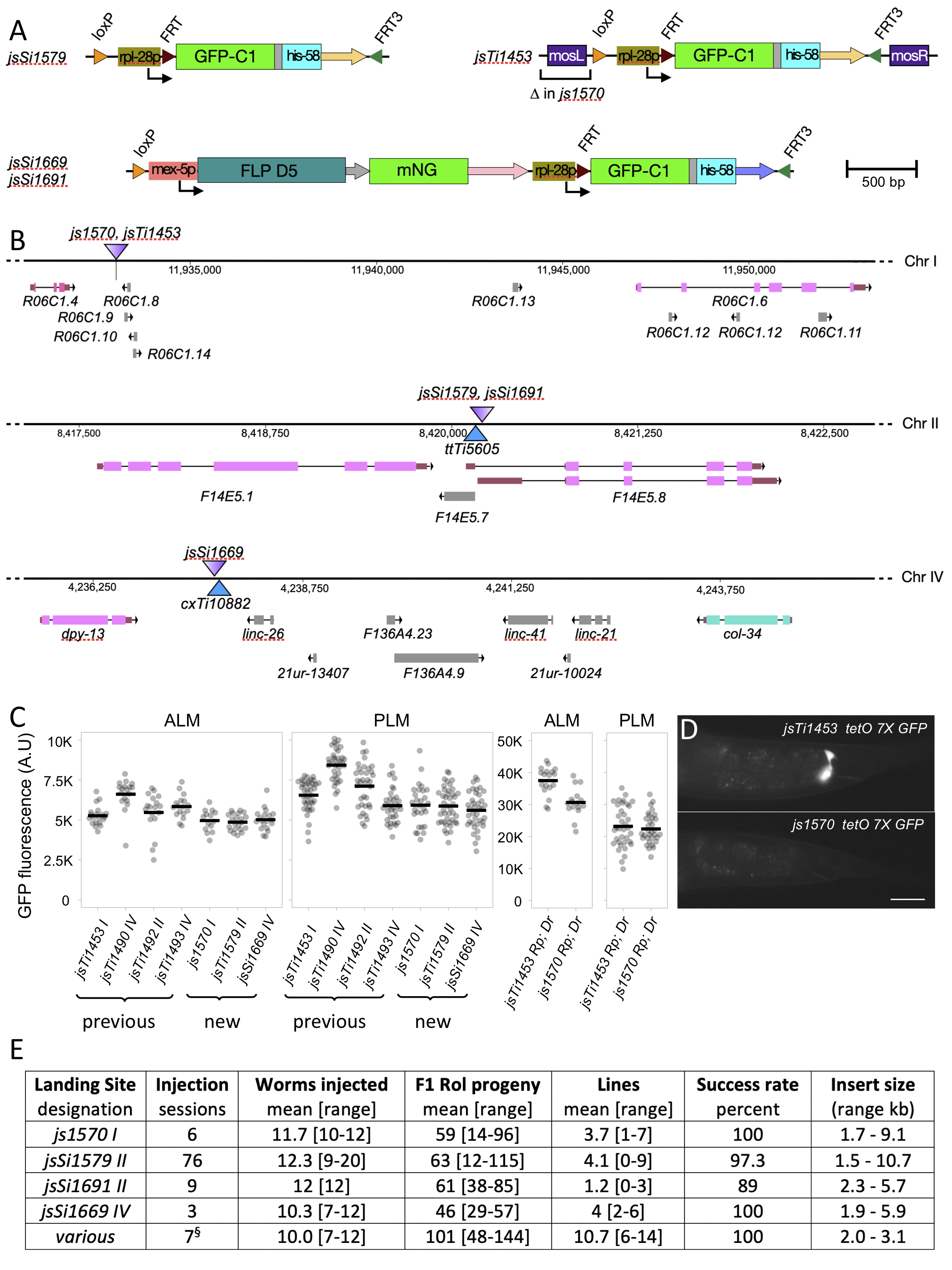
Description
Methods
Reagents
Funding
Author Contributions
- Michael Nonet: Conceptualization, Funding acquisition, Methodology, Formal analysis, Writing - original draft, Writing - review and editing
Reviewed By
Christian Frøkjær-Jensen
Nomenclature Validated By
Anonymous
Database Reference ID: WBPaper00062163
History
- Received: 10/27/2021
- Revision Received: 12/8/2021
- Accepted: 12/15/2021
- Published: 12/16/2021
Copyright
© 2021 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
PubMed Central: PMC8678631
PubMed: 34934910
Cited By
microPublication Biology:ISSN: 2578-9430

